FARSIGHT Framework
Starting Point: We start by assuming that we are given one or more multi-dimensional image(s) I(x,y,z,λ,t), as appropriate. The hardest first step to automated image analysis is image segmentation. We simplify this task using a 'divide and conquer' strategy. We recognize that some of the best-available segmentation algorithms are model based. Usually they require a model describing the expected geometry of biological objects, and a model describing the imaging process and expected defects such as noise and artifacts.
Data Unmixing: Modern microscopes have highly developed spectral imaging hardware and software systems for spectral unmixing. They are very effective at breaking down the image data into a set of non-overlapping channels. In most cases, each channel only contains one type of biological object. This greatly simplifies the task of segmentation by enabling our divide and conquer segmentation strategy (more on this below).
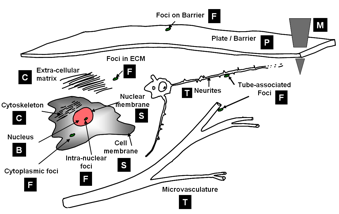
Computational taxonomy of Object Morphologies: We are primarily interested in fluorescence microscopy at the cell and tissue level (0.1 − 1000μm). Within this realm, we have found that, in spite of the variability in biological forms, it is possible to identify a “short list” of frequently occurring morphologies of biological entities – blobs (B), tubes (T), shells (S), foci (F), plates (P), clouds (C), and man-made objects (M). These morphologies are illustrated in the following figure:
Short list of object morphologies frequently observed in fluorescence cell / tissue imagery: B = Blobs; T = Tubes; S = Shells; F = Foci/ punctae; P = Plates; C = Clouds; M = Man-made objects.
Commonly, blobs correspond to cell nuclei, tubes correspond to neuron processes/vasculature, shells correspond to nuclear/cell membranes, foci represent localized molecular concentrations such as mRNA at the site of transcription, cell-cell adhesions and synapses, plates correspond to basal laminae, clouds correspond to cytoplasmic markers, and man-made objects correspond to implanted devices.
Which class should I choose? We require the user to designate what morphological category a given biological object corresponds to. This information specifies a segmentation algorithm from our library of segmentation algorithms. Each algorithm is specialized for the corresponding morphological category. The morphological class of an image object can sometimes be interpreted in more than one manner, suggesting different segmentation algorithm choices. For instance, as foci become larger, it may be advantageous to interpret them as blobs instead. The optimal choice is the one that leads to the lowest segmentation error. Specialized algorithms can achieve higher levels of automation, accuracy, speed, and robustness.
Dealing with an imperfect segmentation: Even the best-available segmentation algorithms have a non-zero error rate. In order to apply them on a large scale to complex problems, it is essential to ensure that: (i) they are run with optimal parameter settings; and (ii) their outputs are inspected and validated before computing the quantitative measurements of eventual interest. With this in mind, the FARSIGHT system will incorporate two innovative software tools. The rapid prototyping system (RPS) is our computer-aided tool for seeking out the best parameter settings, and our edit-based validation system (EVS) is a tool for efficient inspection and corrective editing of segmentations.
Computing Measurements - Round 1: Once the user is satisfied with the segmentation results, he/she can compute diverse measurements (features) of objects in the image and their associations. The diversity of biological investigational objectives implies that different measurements are important for different studies. Nevertheless, the types of measurements of interest fall into two broad categories: (i) intrinsic; and (ii) associative. Intrinsic measurements quantify the features of objects in a single channel. Associative measurements, on the other hand, quantify relationships between objects from one or more channels. We are interested in spatial associations as well as associations over time (e.g., tracking assignments). Most of these features are intended for use by the biologist. In addition to these "end use" features, we also compute some "diagnostic features" whose purpose is to help diagnose segmentation/classification errors.
Object Classification: Using intrinsic and associative features, we are able to classify objects in the image (usually cell types). For this, the FARSIGHT toolkit provides a library of pattern analysis algorithms. In some applications, the analysis ends with object classification. In others, it sets the stage for further analysis, as described below.
Computing Measurements - Round 2: Once all biological objects have been segmented and categorized, a fresh round of quantitative analysis becomes possible. For this, we rely on extensive use of graph theory. This is our Tissue Nets module.
FARSIGHT Image Analysis Pipeline: